Researchers at the University of British Columbia have discovered a key vulnerability across all major variants of the SARS-CoV-2 virus, including the recently emerged BA.1 and BA.2 Omicron subvariants.
The weakness can be targeted by neutralizing antibodies, potentially paving the way for treatments that would be universally effective across variants.
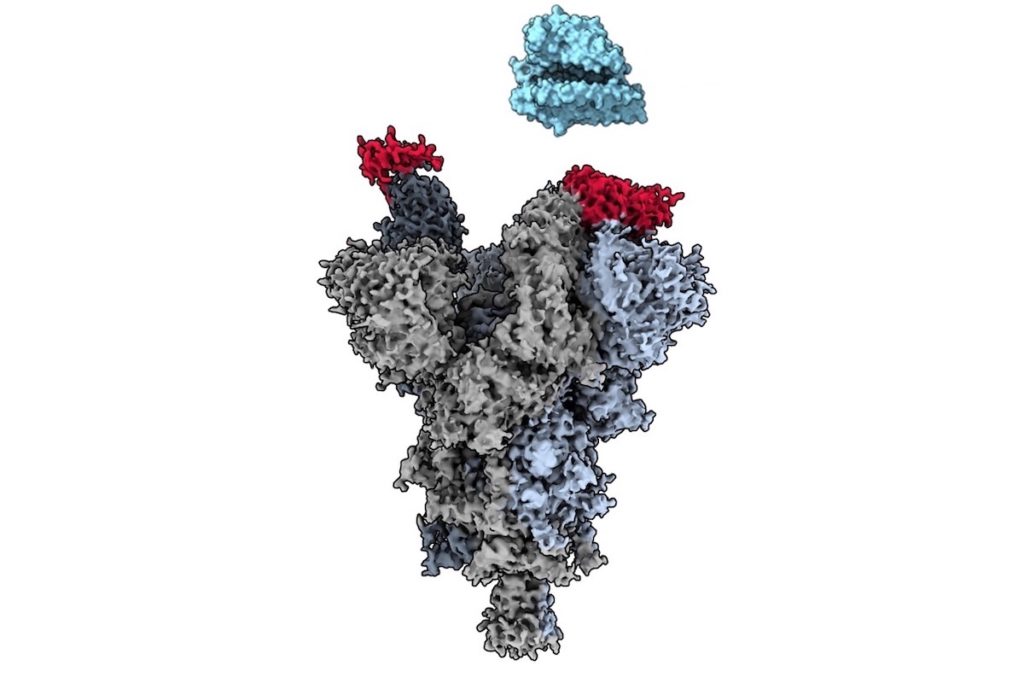
The findings, published today in Nature Communications, use cryo-electron microscopy (cryo-EM) to reveal the atomic-level structure of the vulnerable spot on the virus’ spike protein, known as an epitope. The paper further describes an antibody fragment called VH Ab6 that is able to attach to this site and neutralize each major variant.

Dr. Sriram Subramaniam
“This is a highly adaptable virus that has evolved to evade most existing antibody treatments, as well as much of the immunity conferred by vaccines and natural infection,” says Dr. Sriram Subramaniam (he/him), a professor at UBC’s faculty of medicine and the study’s senior author. “This study reveals a weak spot that is largely unchanged across variants and can be neutralized by an antibody fragment. It sets the stage for the design of pan-variant treatments that could potentially help a lot of vulnerable people.”
Identifying COVID-19 master keys
Antibodies are naturally produced by our bodies to fight infection, but can also be made in a laboratory and administered to patients as a treatment. While several antibody treatments have been developed for COVID-19, their effectiveness has waned in the face of highly-mutated variants like Omicron.
“Antibodies attach to a virus in a very specific manner, like a key going into a lock. But when the virus mutates, the key no longer fits,” says Dr. Subramaniam. “We’ve been looking for master keys — antibodies that continue to neutralize the virus even after extensive mutations.”
The ‘master key’ identified in this new paper is the antibody fragment VH Ab6, which was shown to be effective against the Alpha, Beta, Gamma, Delta, Kappa, Epsilon and Omicron variants. The fragment neutralizes SARS-CoV-2 by attaching to the epitope on the spike protein and blocking the virus from entering human cells.
The discovery is the latest from a longstanding and productive collaboration between Dr. Subramaniam’s team at UBC and colleagues at the University of Pittsburgh, led by Drs. Mitko Dimitrov and Wei Li. The team in Pittsburgh has been screening large antibody libraries and testing their effectiveness against COVID-19, while the UBC team has been using cryo-EM to study the molecular structure and characteristics of the spike protein.
Focusing in on COVID-19’s weak points
The UBC team is world-renowned for its expertise in using cryo-EM to visualize protein-protein and protein-antibody interactions at an atomic resolution. In another paper published earlier this year in Science, they were the first to report the structure of the contact zone between the Omicron spike protein and the human cell receptor ACE2, providing a molecular explanation for Omicron’s enhanced viral fitness.
By mapping the molecular structure of each spike protein, the team has been searching for areas of vulnerability that could inform new treatments.
“The epitope we describe in this paper is mostly removed from the hot spots for mutations, which is why its capabilities are preserved across variants,” says Dr. Subramaniam. “Now that we’ve described the structure of this site in detail, it unlocks a whole new realm of treatment possibilities.”
Dr. Subramaniam says this key vulnerability can now be exploited by drug makers, and because the site is relatively mutation-free, the resulting treatments could be effective against existing—and even future—variants.
“We now have a very clear picture of this vulnerable spot on the virus. We know every interaction the spike protein makes with the antibody at this site. We can work backwards from this, using intelligent design, to develop a slew of antibody treatments,” says Dr. Subramaniam. “Having broadly effective, variant-resistant treatments would be a game changer in the ongoing fight against COVID-19.”
Interview language(s): English